What is a tree sequence?
A succinct tree sequence, or tree sequence for short, represents the relationships between a set of DNA sequences (also known as an ARG). Tree sequences can be used to store genetic data efficiently, and enable powerful analysis of millions of whole genomes at a time. They can be created by simulation or by inferring relationships from genetic variation.
For a fully functional taster of what tree sequences are and how they work, try our interactive workbook.
Tree sequences provide:
Browse tutorials, publications and videos:
Simplification
28 April, 2026
The simplify() method provides several powerful and flexible ways
to modify a tree sequence, and forms the basis of efficient forward-time
simulation of genetic genealogies (“prospective ARGs”). Indeed, it has
been described as the “swiss-army knife” of tskit methods. This tutorial
provides a gentle introduction to simplification, and is accompanied by a
(partially complete) “Advanced simplification” tutorial for experts.
On ARGs, pedigrees, and genetic relatedness matrices
02 December, 2025
GENETICS (2025) Lehmann et al.
This paper discusses branch relatedness in the context of tree sequences,
and introduces a method to use the tree sequence encoding to efficiently
multiply the genetic relatedness matrix (GRM) by a vector, even for huge
sample sizes. This is used to implement a fast way of performing
principle component analysis (PCA) on ARGs and other tree sequences
(now implemented as ts.pca() in tskit)
A forest is more than its trees: haplotypes and ancestral recombination graphs
02 December, 2025
GENETICS (2025) Fritze et al.
This paper discusses how non-coalescent segments of nodes in an
ancestral recombination graph (ARG) are important, shows how
to infer them, and discusses why this can make tree sequence
encodings of ARGs more efficient. The functionality is
implemented in the tskit ts.extend_haplotypes() method.
The paper also describes haplotype-aware methods to compare
tree sequences, implemented in the tscompare library.
Untangling ancestral recombination graphs to infer complex demographic scenarios
12 November, 2024
Nate Pope at Phyloseminar.org
A talk at phyloseminar.org, that explains calculation and use of empirical pair and trio coalescence rate distributions from ARGs. Demonstrates fitting of multipopulation demographic models from inferred coalescent rate distributions, including from inferred ARGs.
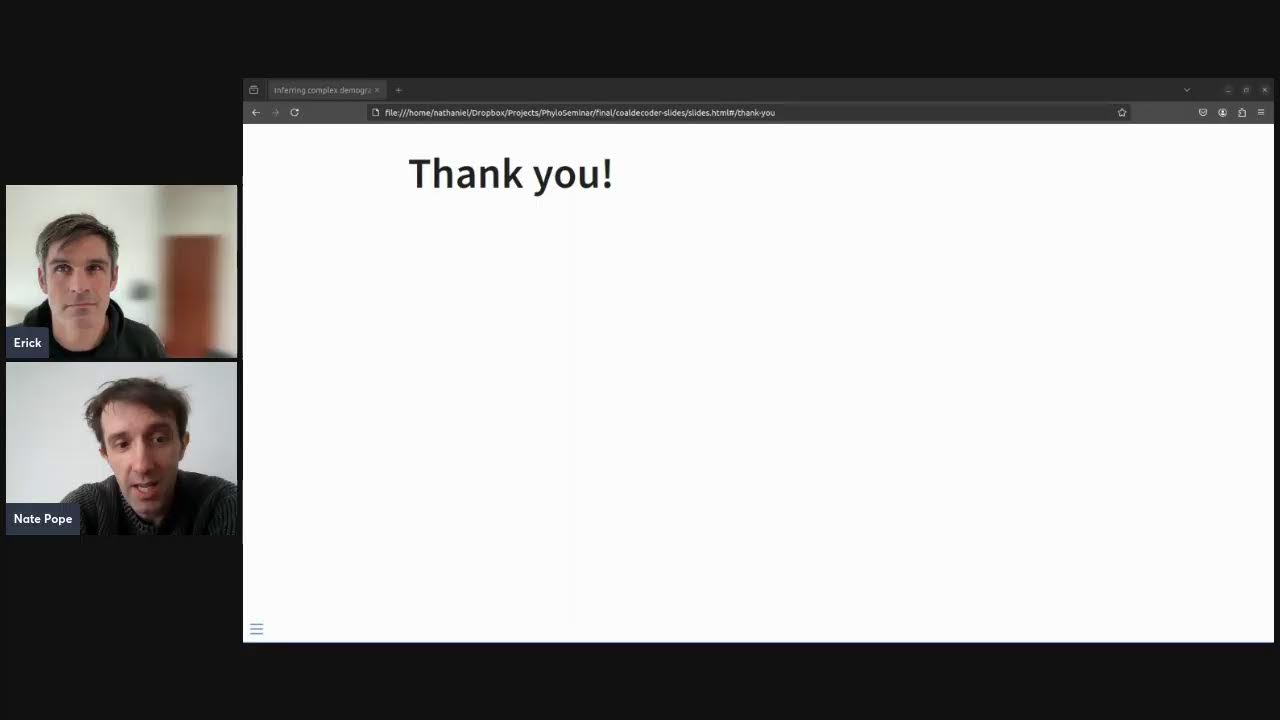
Bayesian inference of Ancestral Recombination Graphs- progress and challenges
15 October, 2024
Yun Deng at Phyloseminar.org
A talk at phyloseminar.org, introducing SINGER, an MCMC sampler for ARGs that scales to hundreds of samples, and applying it to demographic inference an identifying selection.
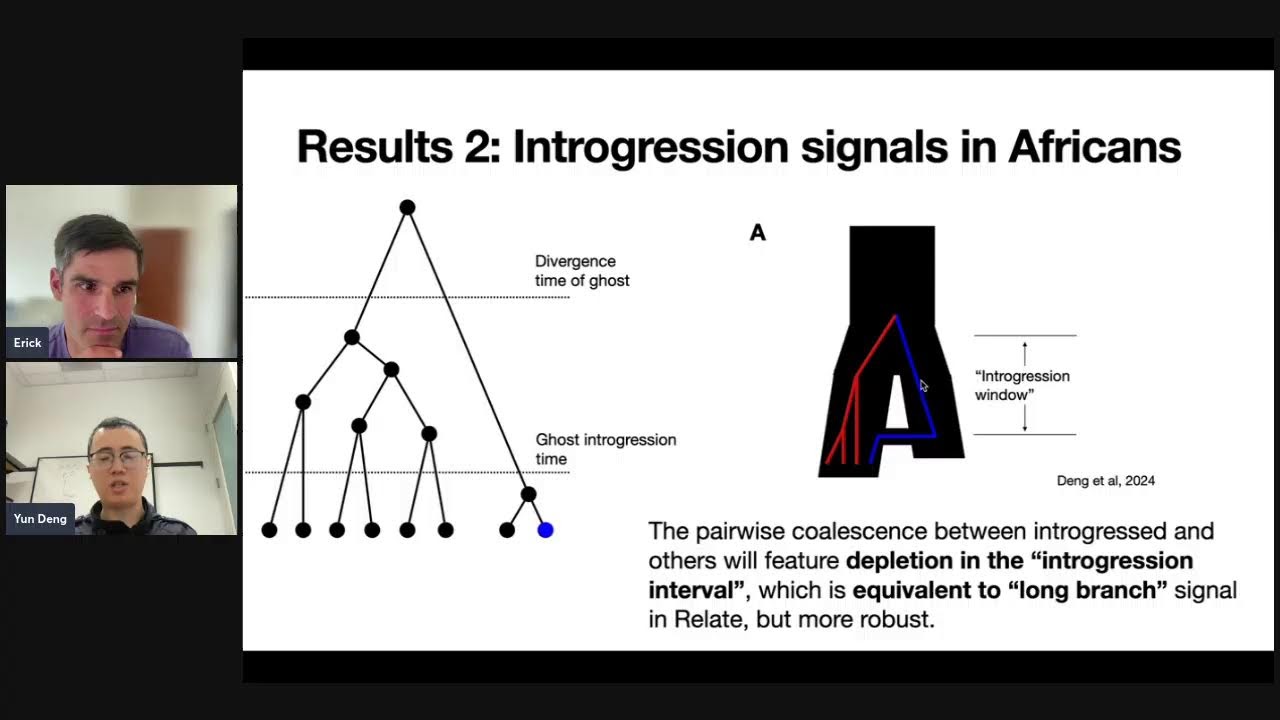
Scalable approaches to inference and analysis of genome-wide genealogies
10 September, 2024
Arni Gunnarsson at Phyloseminar.org
A talk at phyloseminar.org, discussing inferring ARGs using Threads, which uses an operation called “threading” and scales to biobank-scale samples. Shows some applications to imputation and GWAS.
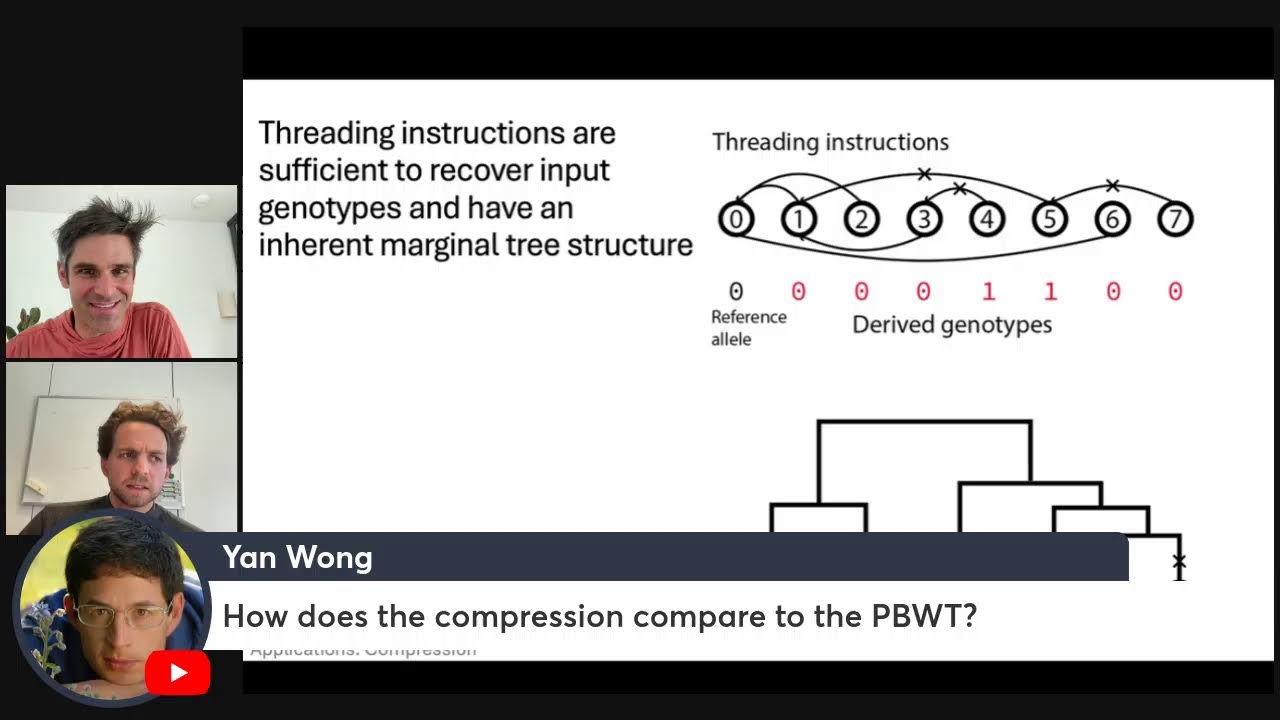
A general and efficient representation of ancestral recombination graphs
06 September, 2024
GENETICS (2024) Wong et al.
This paper recounts recent developments in competing representations of Ancestral Recombination Graphs. It presents a simple formalism that defines an ARG in terms of specific genomes and their intervals of genetic inheritance, showing how this generalizes to encompasses the outputs of recent inference methods. Also see the tskit news item linking to a layman’s summary.
tstrait : a quantitative trait simulator for ancestral recombination graphs
25 May, 2024
Bioinformatics (2024) Tagami et al.
doi: 10.1093/bioinformatics/btae334
This describes a new tskit-based simulator that maps genotypes to phenotypes efficiently for large numbers
of genomes. The new software, tstrait, takes an existing tree sequence containing mutations,
and simulates a quantitative phenotypic trait whose value is partially determined by a selected
set of causal mutations in the tree sequence. Also see the
tskit news item about the associated software release.
The era of the ARG: An introduction to ancestral recombination graphs and their significance in empirical evolutionary genomics
18 January, 2024
PLOS Genetics (2024) Lewanski et al.
doi: 10.1371/journal.pgen.1011110
Although not written by anyone from the core tskit development community, this review gives a good general introduction to Ancestral Recombination Graphs, and discusses how tree sequences can be used to represent them.
link-ancestors: fast simulation of local ancestry with tree sequence software
20 November, 2023
Bioinformatics Advances (2023) Tsambos et al.
Describes an efficient algorithm for calculating “local ancestry” from tree sequences simulated by e.g. msprime or SLiM. This takes a tree sequence and a set of ancestors, and returns a record of the corresponding local ancestry.
ARGs as tree sequences
10 November, 2023
The tskit library provides a convenient way to encode and work with
ancestral recombination graphs (ARGs). This tutorial introduces the
concept of “full ARGs” in tskit, the link between these ARGs and
more simplified graphical representations, and how to use tskit
to work with inheritance graphs in general.
On the genes, genealogies, and geographies of Quebec
25 May, 2023
Science (2023) Anderson-Trocmé et al.
This paper uses a new feature implemented in msprime to simulate genetic transmission through a known pedigree. In this case, a human pedigree based on 4 million Canadian genealogical records was used to highlight the multiple impacts of geography on human migration. The paper is also associated with a freely available simulated whole-genome dataset of almost 1.5 million individuals.
The ARG revolution in population and statistical genetics
30 September, 2022
Wilder Wohns at SMBE Everywhere
In this video, Wilder Wohns gives examples of how ancestral recombination graphs (ARGs)
stored in tskit format have the potential to revolutionise population and statistical
genetics. In particular, inferred tree sequences can be used for inferring geographical
locations of human ancestry, and can be used to improve the speed and accuracy of genetic
association analysis.

A unified genealogy of modern and ancient genomes
25 February, 2022
Science (2022) Wohns et al.
This paper describes using tskit, tsinfer and tsdate to create
a unified tree sequence of 3601 modern and 8 ancient human genome
sequences compiled from eight datasets. Then estimates
of ancestor geographic location are introduced that
recapitulate key features of human history.
Efficient ancestry and mutation simulation with msprime 1.0
13 December, 2021
Genetics (2021) Baumdicker et al.
The accompanying paper to the msprime 1.0 release, summarising its features
and performance and discussing its development model.
Tskit Terminology and Concepts
21 June, 2021
This tutorial serves as an introduction to the terminology and concepts in tskit, and its underlying data structures.
Getting started with tskit
21 June, 2021
You’ve run some simulations or inference methods, and you now have a TreeSequence object; what now? This tutorial is aimed users who are new to tskit and would like to get some basic tasks completed.
Analysing Tree Sequences
21 June, 2021
This tutorial aims to give a quick overview of how the tskit statistics APIs work and how to use them effectively.
Tables and Editing
19 June, 2021
The underlying representation of a tree sequence in tskit is a set of tables. This tutorial shows how to access and
manipulate these tables.
Analysing Trees
19 June, 2021
tskit provides single tree traversals, algorithms and phylogenetic statistics, of which this tutorial gives an overview.
Working with Metadata
16 June, 2021
This tutorial gives an overview of tskit’s metadata system. This allows arbitrary, documented metadata to be attached to
entities in tree sequences.
Do you really need mutations?
06 June, 2021
In tree sequences, the genetic genealogy exists independently of the mutations that generate genetic variation, and often we are primarily interested in genetic variation because of what it can tell us about those genealogies. This tutorial aims to illustrate when we can leave mutations and genetic variation aside and study the genealogies directly.
Visualization
29 May, 2021
It is often helpful to visualize a single tree — or multiple trees along a tree sequence — together with
sites and mutations. tskit provides functions to do this, outputting either plain ascii or unicode text,
or the more flexible Scalable Vector Graphics (SVG) format. This tutorial illustrates various examples.
Tskit and R
11 May, 2021
To interface with tskit in R, we can use the reticulate R package, which lets you call
Python functions within an R session. In this short tutorial, we’ll go through a couple of
examples to show you how to get started.
Completing forwards simulations
20 January, 2021
In this tutorial we show how to combine the best of both forwards and backwards simulation approaches by simulating
the recent past using a forwards-time simulator and then complete the simulation of the ancient past using msprime.
msprime tutorials
19 January, 2021
A set of tutorials for msprime. Covering demography, bottlenecks and introgression.
Efficiently Summarizing Relationships in Large Samples: A General Duality Between Statistics of Genealogies and Genomes
01 July, 2020
Genetics (2020) Ralph et al.
doi: 10.1534/genetics.120.303253
This paper shows that we can think about any statistic that works on sequence data in an equivalent (and more powerful) way in terms of the underlying trees, and that we can compute these statistics very efficiently. Read this paper if you would like more technical details on how the underlying data structures work and an introduction to incremental tree sequence algorithms.
Population genetics using inferred genealogies and tree sequences
17 June, 2020
Leo Speidel at Phyloseminar.org
A talk at phyloseminar.org, that reviews the quickly gorwing set of tools that extract information about evolutionary past from ARGs, focused on Relate and tsinfer.
Tree sequences and inference
22 May, 2020
Yan Wong at Phyloseminar.org
A walk through of the workings of the tsinfer algorithm,
a rapid way to infer tskit tree sequences from existing genetic variation data.
A talk at phyloseminar.org.

Tree sequence fundamentals
08 April, 2020
Wilder Wohns at Phyloseminar.org
A talk at phyloseminar.org, that gives an overview of tskit and the related ecosystem (as of 2020).
Inferring whole-genome histories in large population datasets
02 September, 2019
Nature Genetics (2019) Kelleher et al.
doi: 10.1038/s41588-019-0483-y
Start here if you’re new to tree sequences. This paper introduces tsinfer, the method to infer tree sequence topologies from genetic variation data. Please see the preprint if you cannot access the Nature Genetics paper.
Succinct tree sequences for megasample genomics (47:03)
26 April, 2019
Jerome Kelleher at MIA

Tree‐sequence recording in SLiM opens new horizons for forward‐time simulation of whole genomes
22 November, 2018
Molecular Ecology Resources (2019) Haller et al.
Continuing on from the 2018 PLOS Computational Biology paper, we discuss here how the tree sequence recording method was implemented in the powerful SLiM simulator. We show how some simulations are orders of magnitude more efficient and examples of the new possibilities that keeping a full record of the genetic ancestry makes available.
Efficient pedigree recording for fast population genetics simulation
01 November, 2018
PLOS Computational Biology (2018) Kelleher et al.
doi: 10.1371/journal.pcbi.1006581
Forwards-in-time simulations are very flexible but also usually very CPU intensive. This paper shows how we used tree sequences to make forwards-in-time simulations both more efficient and even more flexible.
Simulating, storing & processing genetic variation data for millions of samples
26 April, 2017
Jerome Kelleher at MIA

Efficient Coalescent Simulation and Genealogical Analysis for Large Sample Sizes
04 April, 2016
PLOS Computational Biology (2016) Kelleher et al.
doi: 10.1371/journal.pcbi.1004842
This is where the tree sequence data structure was first described. Here we introduce the msprime coalescent simulator and the core algorithms and data structures that would later be separated out into tskit. Read this paper if you would like to find out more about coalescent simulation, or to understand the core tree sequence algorithms and theoretical results. Note: much of the terminology has been updated since this original publication as the models were generalised.
